About Us
Our Mission
Driving innovation, building industry, empowering change
Our Mission
Accelerating the commercialisation of engineering biology
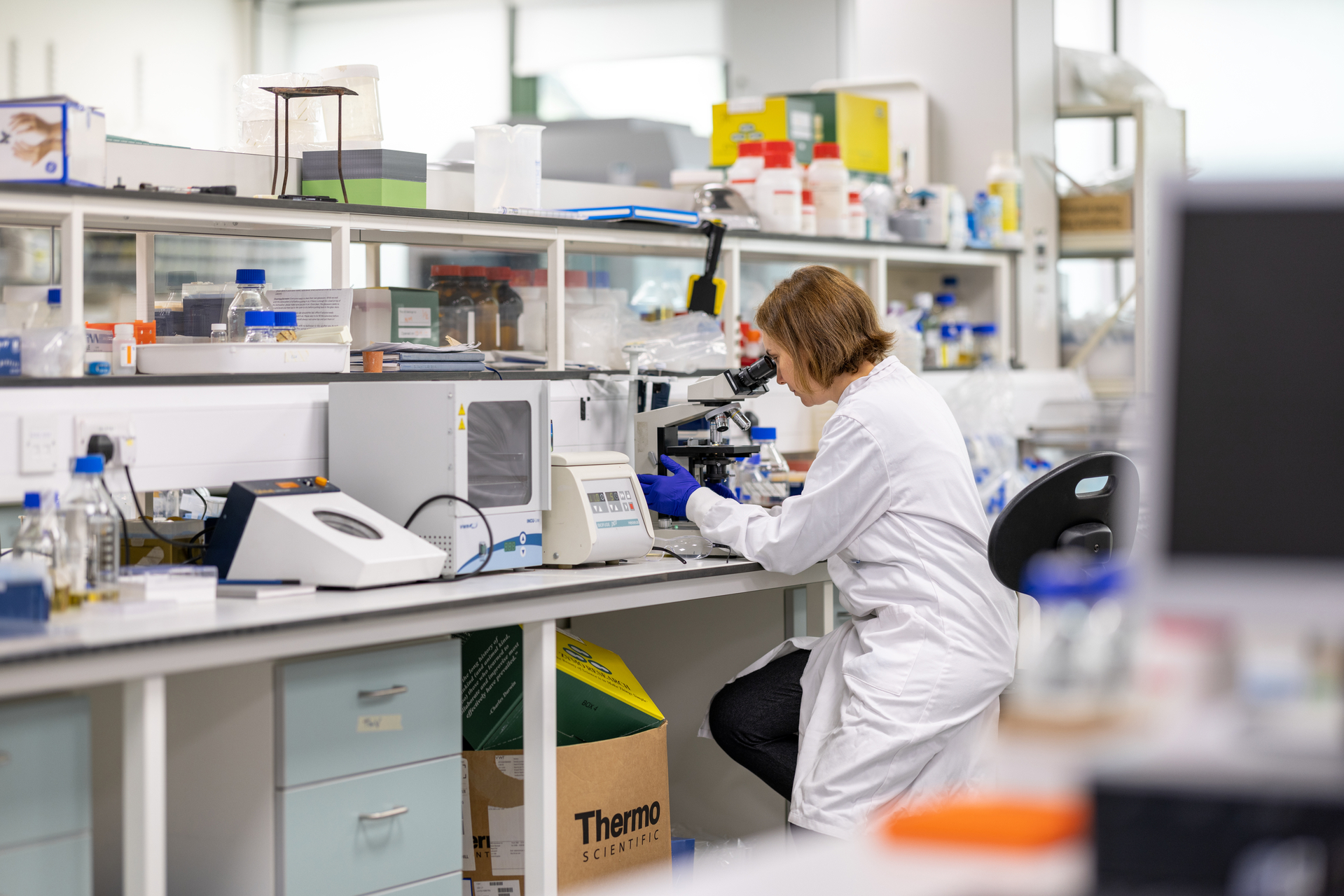

At SynbiCITE, we are dedicated to accelerating the commercialisation of engineering biology to create transformative solutions for the UK economy and beyond.
By uniting academia, industry, and policy, we deliver substantial and sustained benefits to the UK bioeconomy.
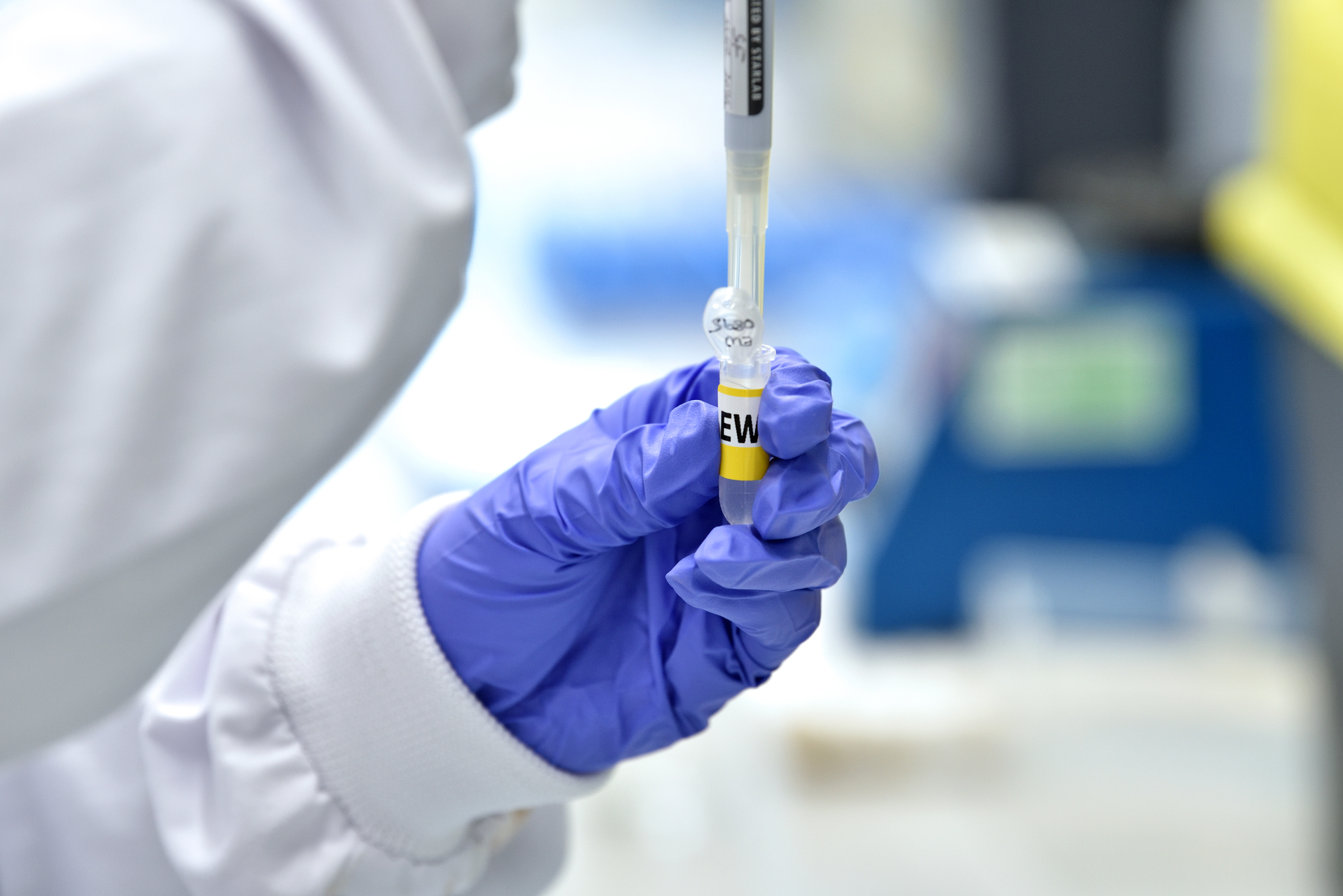

Our Mission
Evolving our industry
As the UK’s national industrial translation centre for engineering biology, our mission is to act as a catalyst for this rapidly evolving industry by:
- Promoting the adoption of engineering biology across industries and acceleratng industrial translation and commercialisation of engineering biology solutions.
- Supporting and scaling start-ups and SMEs to foster innovation and growth by providing business education and a supportive business environment.
- Bridging the gap between research and industrial application by building an expert workforce.
- Supporting and developing the UK’s basic research infrastructure through national and international partnerships.
Responsible Innovation
Our commitment to responsible innovation
We are committed to responsible innovation that prioritises ethical, societal, and environmental considerations. Engineering biology has the power to address critical global challenges.
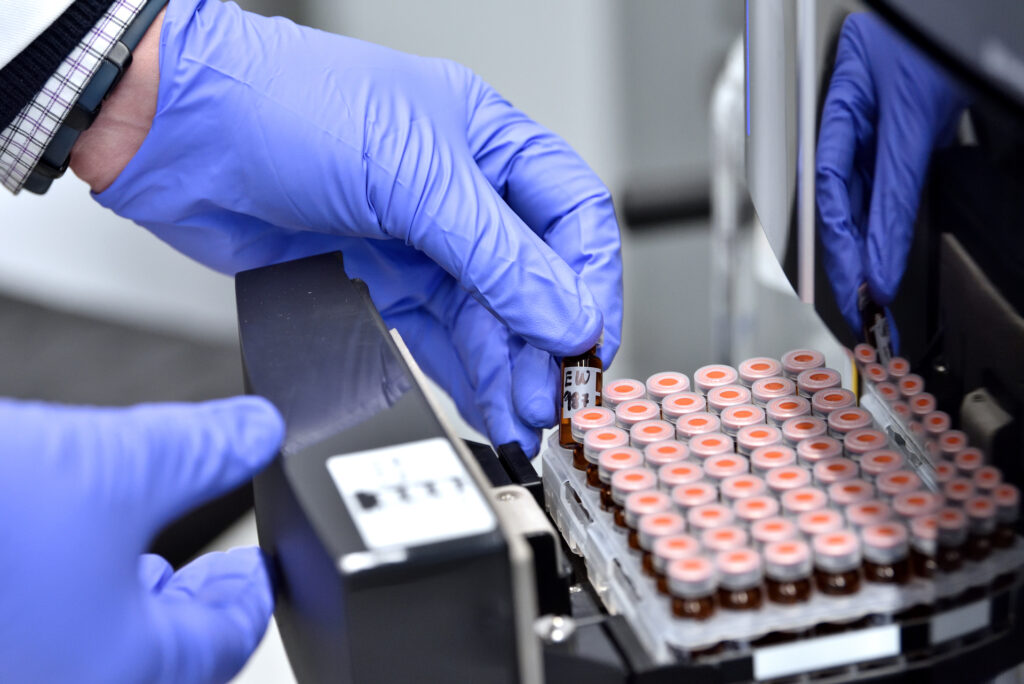

The Future
Our vision for the future
With a commitment of £2 billion over the next decade, the UK is prioritising engineering biology as one of five critical technologies shaping the future. SynbiCITE is proud to lead this charge, creating opportunities for economic growth, sustainability, and technological advancement.
Together, we are building a future where engineering biology transforms industries, strengthens the UK’s innovation ecosystem, and empowers global progress.
Case Studies
Success stories from across our ecosystem
Help us build a sustainable, bio-based future
Get in touch with our team